The burden of drug-resistant infections continues to threaten our arsenal of antibiotics. Despite the rise in genomic sequencing of clinical isolates, there is a lack

Rooney et al. 2022 Jun 1;e0002222. Short-read sequencing can provide detection of multiple genomic determinants of antimicrobial resistance from single bacterial genomes and metagenomic samples.

Kim et al. Clin Microbiol Rev. 2022 May 25;e0017921. Antimicrobial resistance (AMR) is a global health crisis that poses a great threat to modern medicine.

Enabling genomic island prediction and comparison in multiple genomes to investigate bacterial evolution and outbreaks
Bertelli et al. Microb Genom. 2022 May;8(5). doi: 10.1099/mgen.0.000818. Outbreaks of virulent and/or drug-resistant bacteria have a significant impact on human health and major economic

May 1st is our annual staff roll-over for the Comprehensive Antibiotic Resistance Database (CARD). We say farewell to Romeo Chalil, who has been CARD Junior

Broadfield et al. Molecular Metabolism 2022. Apr 20;101498. Type 2 diabetes and obesity increase the risk of developing colorectal cancer (CRC). The first-line type 2
Another (mostly) online year, but thanks to our 2021-2022 BiomedDC 4A15, Biochem 4T15, and HthSci 4R09 thesis students for all of their hard work! Thanks

Sanderson et al. 2022. bioRxiv 10.1101/2022.04.11.487771. Enterococcus faecium is a ubiquitous opportunistic pathogen that is exhibiting increasing levels of antimicrobial resistance (AMR). Many of the

Karyn Mukiri – Predicting the Total Resistome Mugdha Dave – Creation and standardization of automated bacterial pathogen genomic sequencing reporting through informatics

Emma J. Griffiths et al. Gigascience. 2022 Feb 16;11:giac003. Background: The Public Health Alliance for Genomic Epidemiology (PHA4GE) (https://pha4ge.org) is a global coalition that is

Brian Alcock has received a Gradaute Student Association Award for Contributions by Non-Academic Staff! Brian works as CARD’s Lead Curator but also has trained countless

The McArthur lab welcomes Madeline McCarthy, a recent MSc graduate of VIDO / USask, as she join us for her PhD studies in the McMaster

Kara K Tsang, Andrew Latchman, Nishma Singhal, Giuliana Federici, Sandra Russell, Denise Irwin, Robyn Stevens, Andrew G McArthur, & Sarah Khan BMC Med Educ. 2021

Emily Panousis wins the Michael Kamin Hart Memorial Undergraduate Scholarship for her work on SARS-CoV-2 surveillance!
More information about IIDR Trainee Day and Michael Kamin Hart here.

September is here and that means new undergraduate thesis students! Emily Panousis & Gene Liang will start their COVID-19 delayed undergraduate thesis, joined by summer

Two current lab members are changing their roles, with Mugdha Dave starting her MSc studies and Dr. Sheridan Baker transitioning from CanCOGeN work to a

Today we say goodbye to William Huynh, who first joined us as an undergraduate thesis student and then stayed on to work as the Comprehensive

Wang, B., E.E. Tsakiridis, S. Zhang, A. Llanos, E.M. Desjardins, J.M. Yabut, A.E. Green, E.A. Day, B.K. Smith, J.S.V. Lally, J. Wu, A.R. Raphenya, K.A.
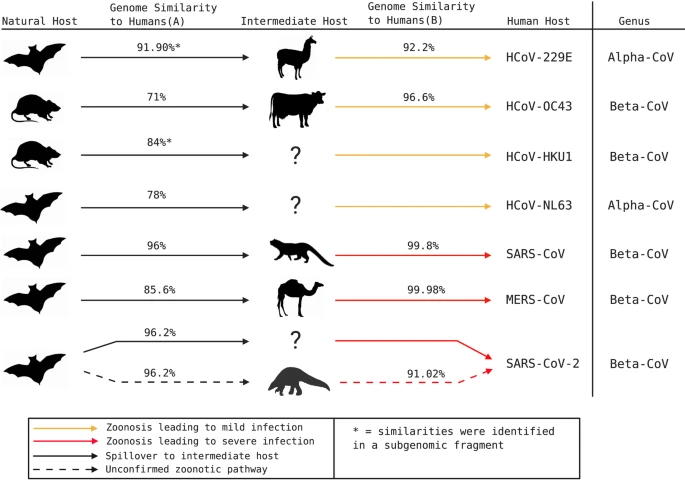
Jalen Singh, Pranav Pandit, Andrew G McArthur, Arinjay Banerjee, Karen Mossman Virology Journal. 2021 Aug 13;18(1):166. The emergence of a novel coronavirus, severe acute respiratory

Jalees Nasir awarded 2021 Fred & Helen Knight Enrichment Award!
Congratulations to PhD student Jalees Nasir, who has been awarded the prestigious 2021 Fred & Helen Knight Enrichment Award, which recognizes exceptional promise in early
