Adam S Komorowski, Xena X Li, Eva Piessens, Andrew G McArthur, & Ameen Patel Journal of the Association of Medical Microbiology and Infectious Disease Canada,
Category: publications

Shehryar Ahmad, Kara L Tsang, Kartik Sachar, Dennis Quentin, Tahmid M Tashin, Nathan P Bullen, Stefan Raunser, Andrew G McArthur, Gerd Prehna, & John C

Sarah Khan, Kara K Tsang, Dominik Mertz, Myrna Dolovich, Marcel Tunks, Catherine Demers, Kelly Hassall, Neil Maharaj, Karen Margallo, Maureen Cividino, Zain Chagla, & MyLinh

Kara K Tsang, Dominik Mertz, Cindy O’Neill, & Sarah Khan Infect Control Hosp Epidemiol. 2020 Nov 4;1-7. As the COVID-19 pandemic evolved, infection prevention and

Workenhe, S.T., A. Nguyen, D. Bakhshinyan, J. Wei, D.N. Hare, K.L. MacNeill, Y. Wan, A. Oberst, J.L. Bramson, J.A. Nasir, S.K. Singh, A.G. McArthur, &
Arinjay Banerjee, Andrew C Doxey, Benjamin J-M Tremblay, Michael J Mansfield, Sonu Subudhi, Jeremy A Hirota, Matthew S Miller, Andrew G McArthur, Samira Mubareka, &

Maya A Farha, Craig MacNair, Lindsey A Carfrae, Sara S El Zahed, Michael Joseph Ellis, Hiu-Ki R Tran, Andrew McArthur, & Eric D Brown ACS

A Comparison of Whole Genome Sequencing of SARS-CoV-2 Using Amplicon-Based Sequencing, Random Hexamers, and Bait Capture
Jalees A. Nasir, Robert A. Kozak, Patryk Aftanas, Amogelang R. Raphenya, Kendrick M. Smith, Finlay Maguire, Hassaan Maan, Muhannad Alruwaili, Arinjay Banerjee, Hamza Mbareche, Brian

Emma J. Griffiths, Ruth E. Timme, Andrew J. Page, Nabil-Fareed Alikhan, Dan Fornika, Finlay Maguire, Catarina Inês Mendes, Simon H. Tausch, Allison Black, Thomas R.
Ana T Duggan, Jennifer Klunk, Ashleigh F Porter, Anna N Dhody, Robert Hicks, Geoffrey L Smith, Margaret Humphreys, Andrea M McCollum, Whitni B Davidson, Kimberly

Arinjay Banerjee, Patrick Budylowski, Daniel Richard, Hassaan Maan, Jennifer A. Aguiar, Nader El-Sayes, Michael R. D’Agostino, Benjamin J.-M. Tremblay, Sam Afkhami, Mehran Karimzadeh, Lily Yip,
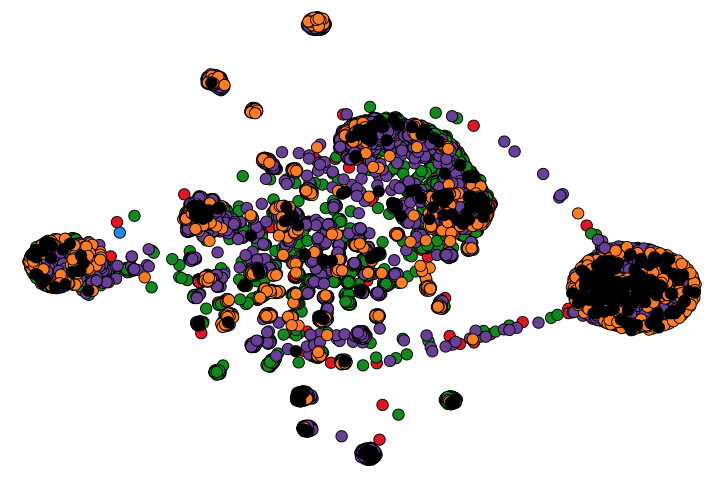

Maan, H., H. Mbareche, A.R. Raphenya, A. Banerjee, J.A. Nasir, R.A. Kozak, N. Knox, S. Mubareka, A.G. McArthur, & B. Wang. The Lancet Digital Health,

Krishna A Srinivasan, Suman K Virdee, Andrew G McArthur Brief Funct Genomics 2020 May 16 [Epub ahead of print] RNA sequencing (RNA-Seq) is a complicated

Chen CY, Clark CG, Langner S, Boyd DA, Bharat A, McCorrister SJ, McArthur AG, Graham MR, Westmacott GR, Van Domselaar G Proteomics Clin Appl. 2019
Day EA, Ford RJ, Smith BK, Mohammadi-Shemirani P, Morrow MR, Gutgesell RM, Lu R, Raphenya AR, Kabiri M, McArthur AG, McInnes N, Hess S, Paré

Ahmad S, Wang B, Walker MD, Tran H-KR, Stogios PJ, Savchenko A, Grant RA, McArthur AG, Laub MT, Whitney JC Nature 2019 Nov 6. [Epub

CARD 2020: antibiotic resistome surveillance with the Comprehensive Antibiotic Resistance Database
Alcock BP, Raphenya AR, Lau TTY, Tsang KK, Bouchard M, Edalatmand A, Huynh W, Nguyen A-LV, Cheng AA, Liu S, Min SY, Miroshnichenko A, Tran

Guitor AK, Raphenya AR, Klunk J, Kuch M, Alcock B, Surette MG, McArthur AG, Poinar HN, Wright GD. Editor’s Pick! Antimicrobial Agents and Chemotherapy 2019 Oct

Waglechner N, McArthur AG, Wright GD. Nat Microbiol. 2019 Aug 12. [Epub ahead of print] Glycopeptide antibiotics are produced by Actinobacteria through biosynthetic gene clusters

PCR detection of mixed and zoonoses Malaria using Plasmodium spp dynein light chain (dlc-tctex) gene
Kariuki MW, Githui E, McArthur AG, Aman RA, Njagi NM, Mwangemi AC, Kamau LW Annual Research & Review in Biology, 32, 1-12. Novel gene targets
