Lu et al. Genome Med. 2025 May 6;17(1):46. Background: Antimicrobial resistant (AMR) pathogens represent urgent threats to human health, and their surveillance is of paramount
Category: big data

Two alumni of the McMaster Biochemistry graduate program have joined the McArthur Lab! Dr. Emily Bordeleau got her PhD from McMaster in 2022 under the

Nirmalarajah et al. BMC Infect Dis. 2025 Jan 28;25(1):132. Background: Drivers of COVID-19 severity are multifactorial and include multidimensional and potentially interacting factors encompassing viral

Jon Stokes (left) and graduate student Autumn Arnold (right) are excited to launch the ESKAPE Model, an all-new AI tool designed to help the global

Mateusz Wlodarski wins the bioMerieux prize for best poster presentation! Arnold, A., A.R. Raphenya, A.G. McArthur, & J.M. Stokes. 2024. The ESKAPE model: AI-guided antibiotic

Tsang et al. Microb Genom. 2024 Oct;10(10). Interpreting the phenotypes of bla SHV alleles in Klebsiella pneumoniae genomes is complex. Whilst all strains are expected

Antimicrobial Resistance – Genomes, Big Data and Emerging Technologies – Wellcome Trust, UK
Mukiri, K.M., B.P. Alcock, & A.G. McArthur. 2024. Increasing the predictive accuracy of the Resistance Gene Identifier by abandoning sole reliance on bitscore. Ta, T.E.,
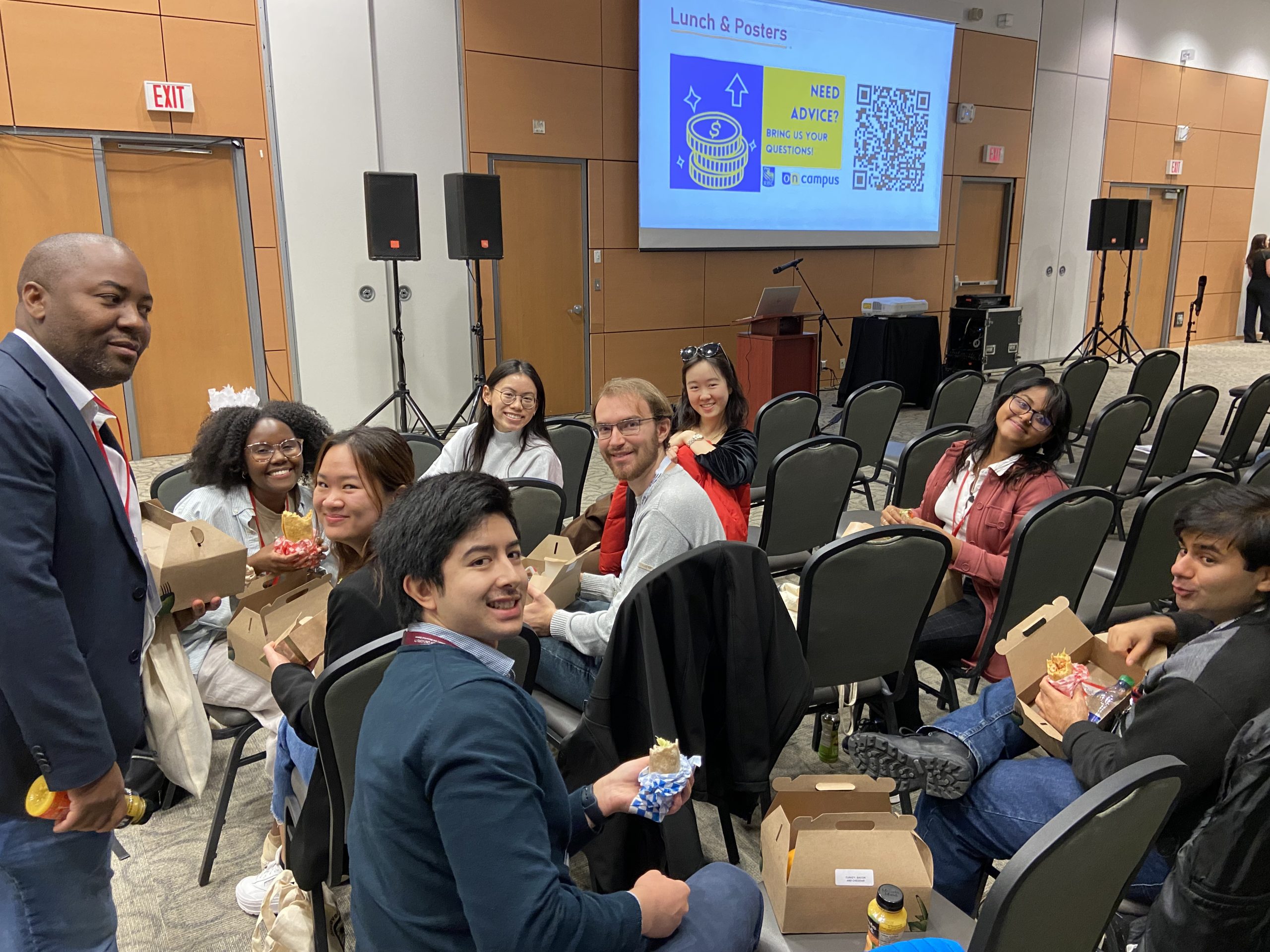

IIDR Trainee Day 2023
COLIN BRUCE – Investigating Fibre Degradation in the Infant Gut Microbiota ; DIRK HACKENBERGER – Was World War 2 Foundational to the Antimicrobial Resistance Crisis?
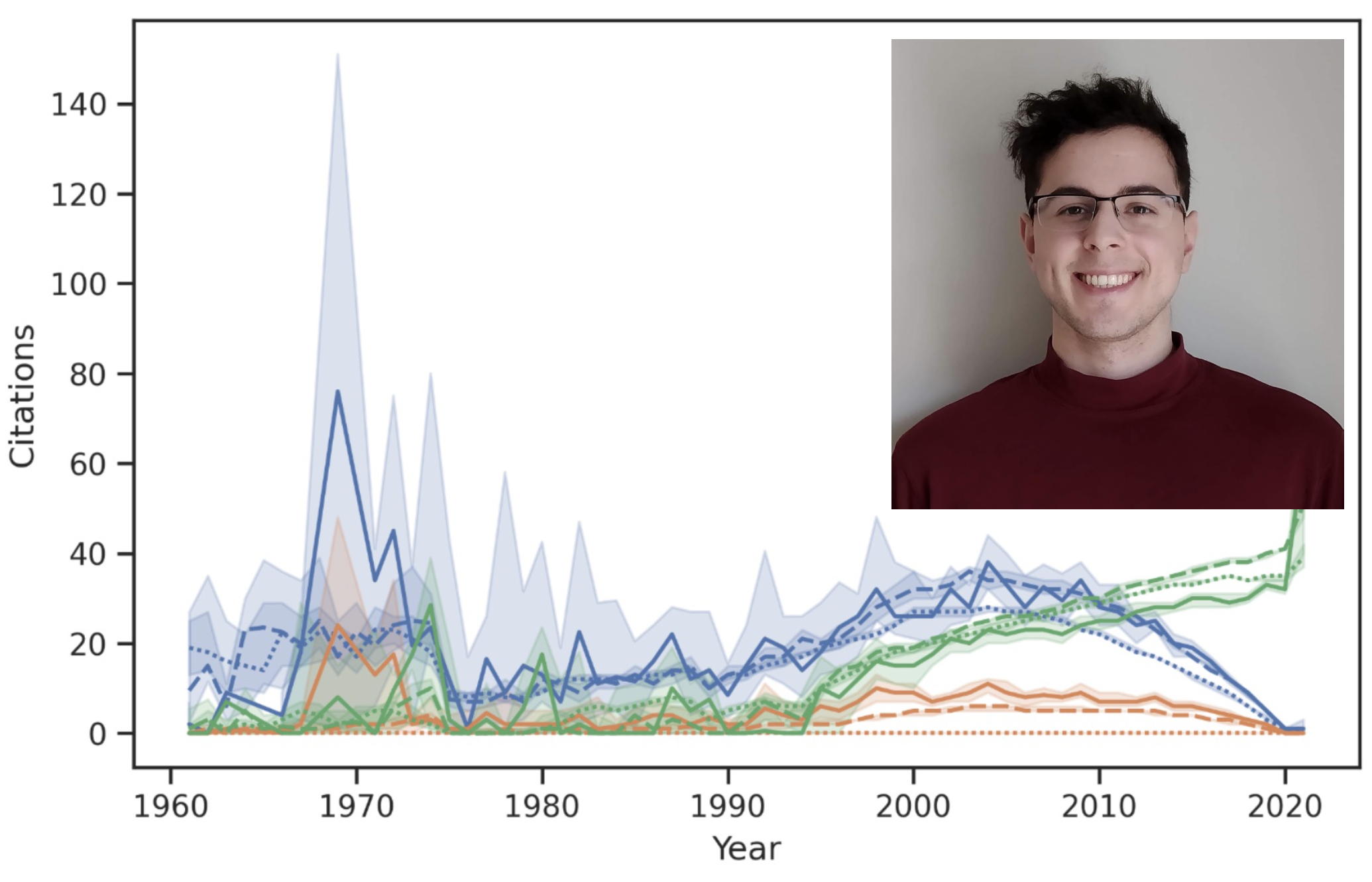
Volunteer starting in 2019, thesis student, summer student, Masters student (2020-2022), and developer, Arman Edalatmand has been a great lab member for over 4 years,

Alcock, B.P., A.R. Raphenya, A. Edalatmand, & A.G. McArthur. 2023. The Comprehensive Antibiotic Resistance Database – curating the global resistome. Oral presentation at the 16th

Arman Edalatmand & Andrew G McArthur. 2023. Database, doi: 10.1093/database/baad023. Scientific literature is published at a rate that makes manual data extraction a highly time-consuming

The Dataharmonizer: a Tool for Faster Data Harmonization, Validation, Aggregation, and Analysis of Pathogen Genomics Contextual Information
Gill et al. 2022. Preprints 10.20944/preprints202206.0335.v1 Pathogen genomics is a critical tool for public health surveillance, infection control, outbreak investigations, as well as research. In

Raphenya et al. Scientific Data volume 9, Article number: 341 (2022). Whole genome sequencing (WGS) is a key tool in identifying and characterising disease-associated bacteria

Kim et al. Clin Microbiol Rev. 2022 May 25;e0017921. Antimicrobial resistance (AMR) is a global health crisis that poses a great threat to modern medicine.

Sanderson et al. 2022. bioRxiv 10.1101/2022.04.11.487771. Enterococcus faecium is a ubiquitous opportunistic pathogen that is exhibiting increasing levels of antimicrobial resistance (AMR). Many of the

Arman Edalatmand has been awarded a 2021 MacData Fellowship from McMaster’s MacData Institute. Working with co-supervisors Drs. Andrew McArthur (Biochemistry & Biomedical Sciences) and Victor

Kara K. Tsang, Finlay Maguire, Haley L. Zubyk, Sommer Chou, Arman Edalatmand, Gerard D. Wright, Robert G. Beiko, & Andrew G. McArthur Microbial Genomics, in
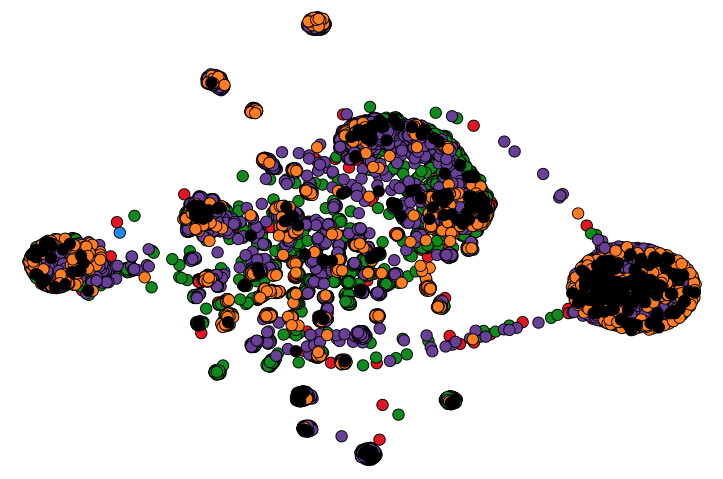

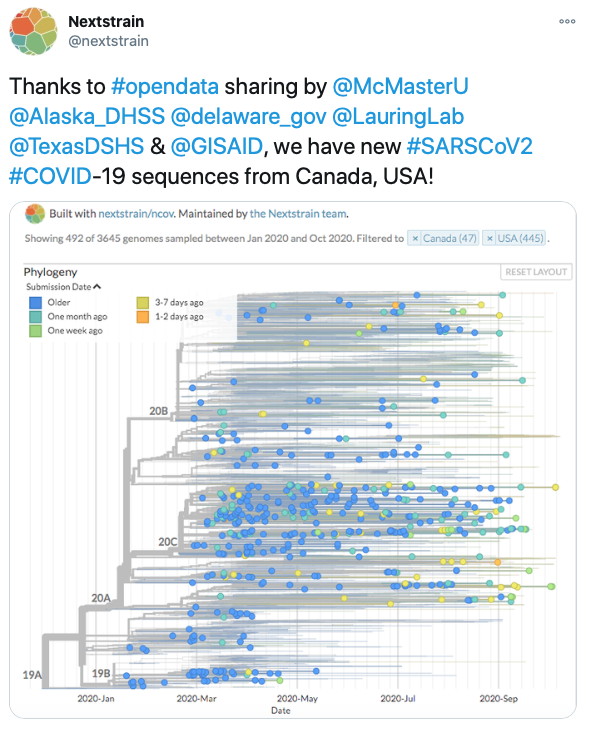
Maan, H., H. Mbareche, A.R. Raphenya, A. Banerjee, J.A. Nasir, R.A. Kozak, N. Knox, S. Mubareka, A.G. McArthur, & B. Wang. The Lancet Digital Health,

CARD 2020: antibiotic resistome surveillance with the Comprehensive Antibiotic Resistance Database
Alcock BP, Raphenya AR, Lau TTY, Tsang KK, Bouchard M, Edalatmand A, Huynh W, Nguyen A-LV, Cheng AA, Liu S, Min SY, Miroshnichenko A, Tran