Kara Tsang passed her graduate transfer exam today, officially moving from the McMaster Biochemistry & Biomedical Sciences Masters program to the Ph.D. program. Kara’s work
Category: genomics
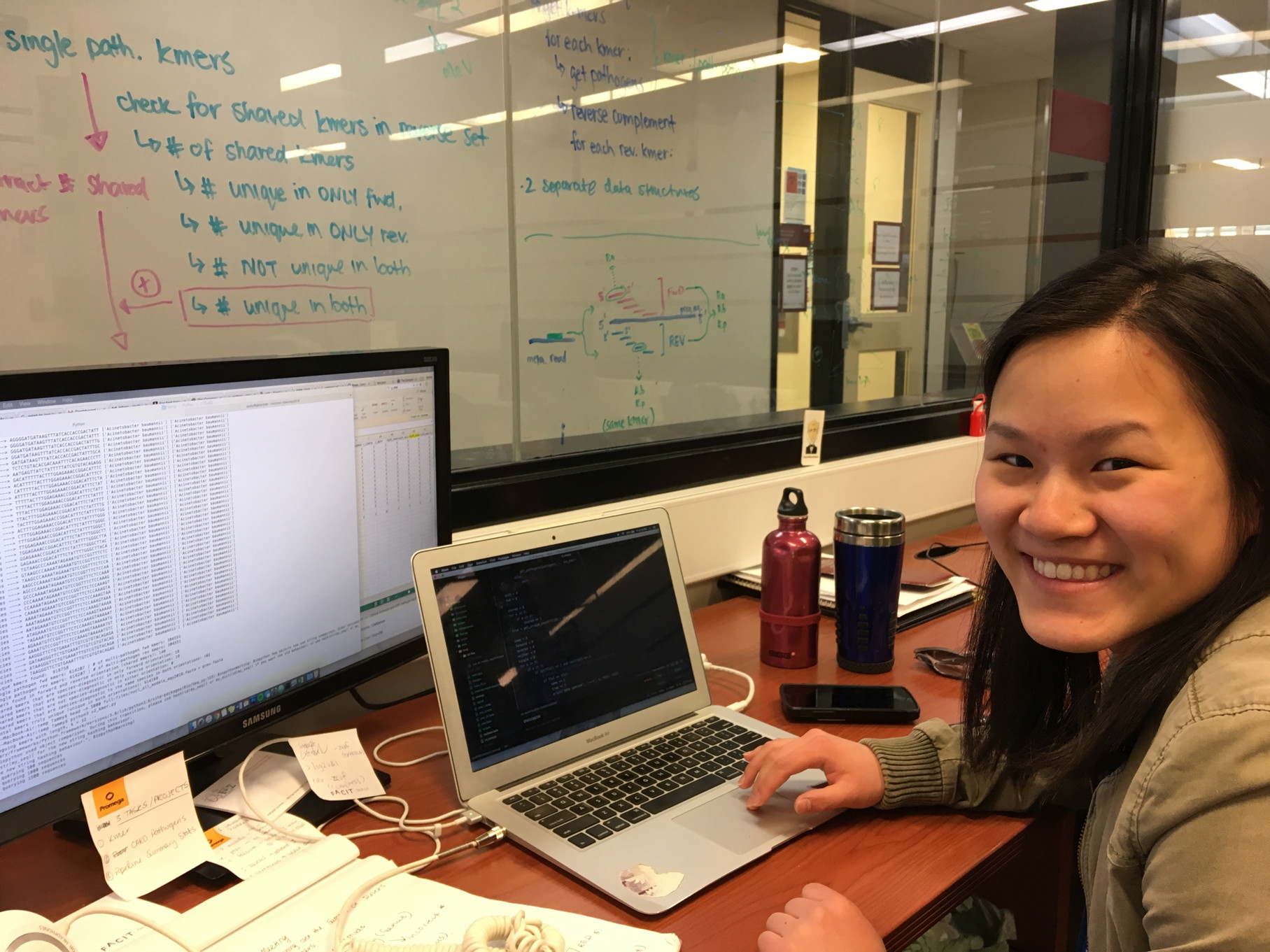

Tammy Lau has been awarded a prestigious Michael G. DeGroote Institute for Infectious Disease Research (IIDR) Summer Student Fellowship for her work on development of k-mer approaches
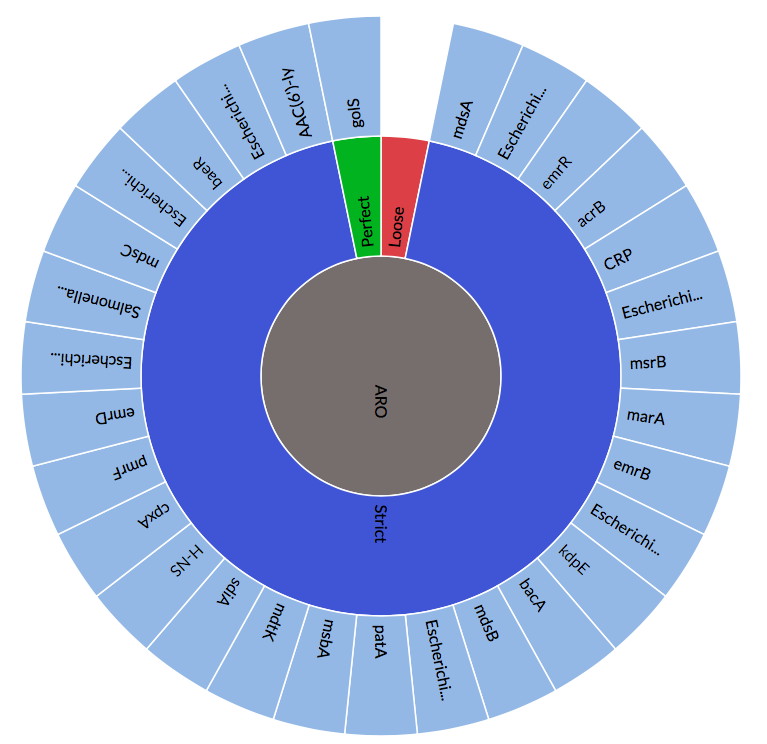
Recent Updates to the Comprehensive Antibiotic Resistance Database
The Comprehensive Antibiotic Resistance Database has been updated, http://card.mcmaster.ca CARD Curation: Addition of HERA, TRU, & ACI beta-lactamases, sul4, and new quinolone efflux pumps. Antibiotic

The Comprehensive Antibiotic Resistance Database: Expanded tools for molecular surveillance of antimicrobial resistance (AMR) in the environment, agriculture, and clinic. A.G. McArthur (PI), G. Van

The McArthur lab and the Comprehensive Antibiotic Resistance Database are proud to join the Canadian Anti-Infective Innovation Network, International Genomic Epidemiology Application Ontology Consortium, and Integrated Rapid
Major Update of the Comprehensive Antibiotic Resistance Database
The Comprehensive Antibiotic Resistance Database has been updated, http://card.mcmaster.ca This February 2018 release is our largest to date and includes new data types, a new

Building upon her successful Biochem 3A03 project, Tammy Lau is staying in the lab for 2017-2018 as part of her Biochem 4T15 Research Thesis. Tammy’s

Pawlowski AC, Wang W, Koteva K, Barton HA, McArthur AG, & Wright GD. Nat Commun. 2016 Dec 8;7:13803. Antibiotic resistance is ancient and widespread in

A cross-national research consortia co-led by McMaster’s Andrew McArthur is receiving two of 16 federal grants to further develop a big data solution to the

McArthur, A.G., B. Jia, A.R. Raphenya, P. Guo, K. Tsang, B. Dave, B. Alcock, B. Lago, N. Waglechner, & G.D. Wright. 2016. The Comprehensive Antibiotic Resistance
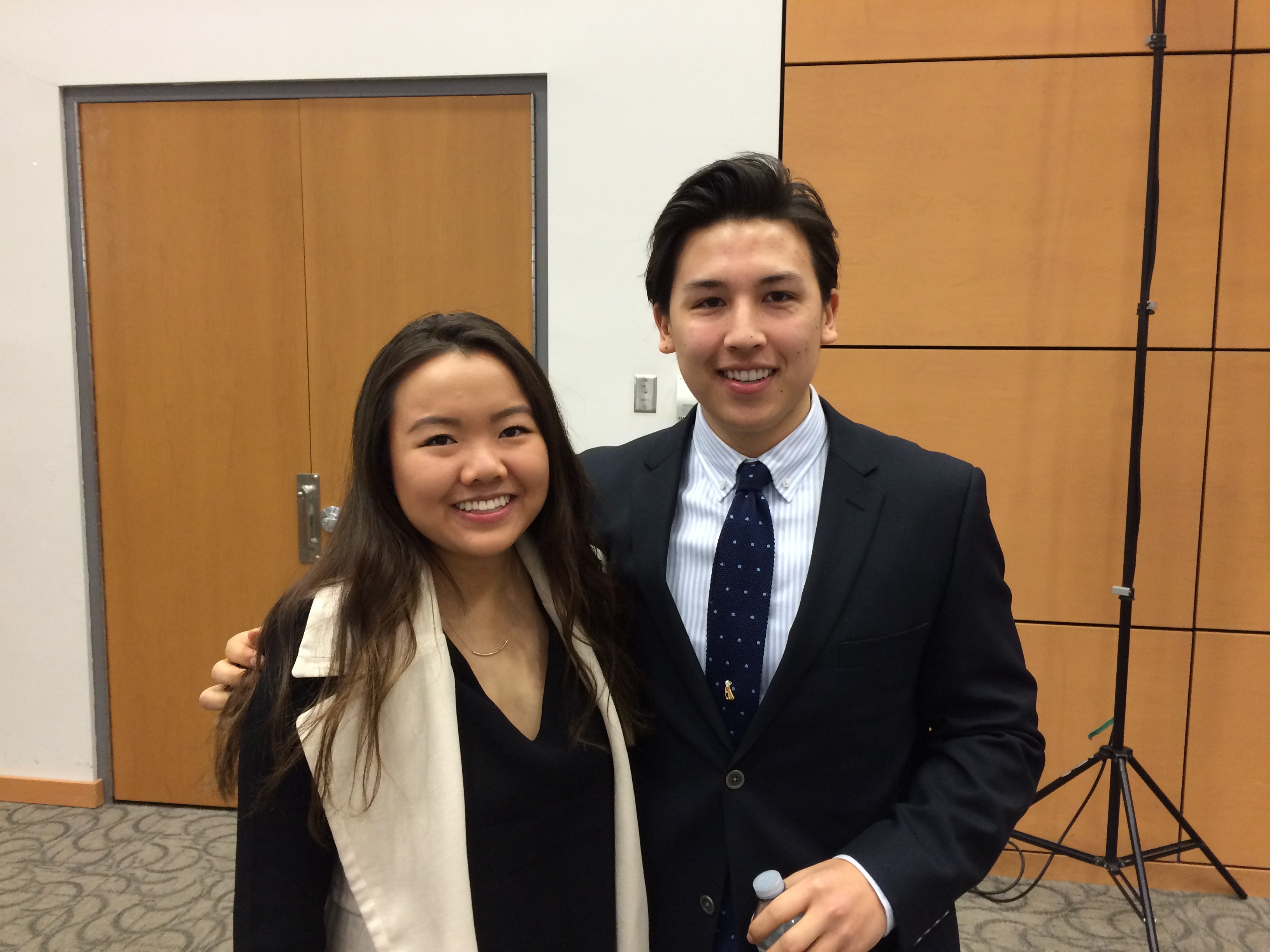

Congratulations to Kara Tsang and Zachary Lin on completion of their Biomedical Discovery and Commercialization (BDC) 4A15 thesis research! Both Kara & Zachary presented their research

Combatting Antibiotic Resistance Using Surveillance – click on the image to watch the 10 minute video. More details here.

The Inaugural Biomedical Discovery & Commercialization (BDC) Symposium
Zachary Lin – Adapting Galaxy bioinformatics to outbreak-associated Clostridium difficile The completion of the human genome project in 2001 sparked the beginning of a sequencing revolution

A new NIH-funded collaboration with Dr. Larissa Williams, Bates College, USA that applies transcriptomic approaches to examination of the role Nfe2 in oxidative stress response during
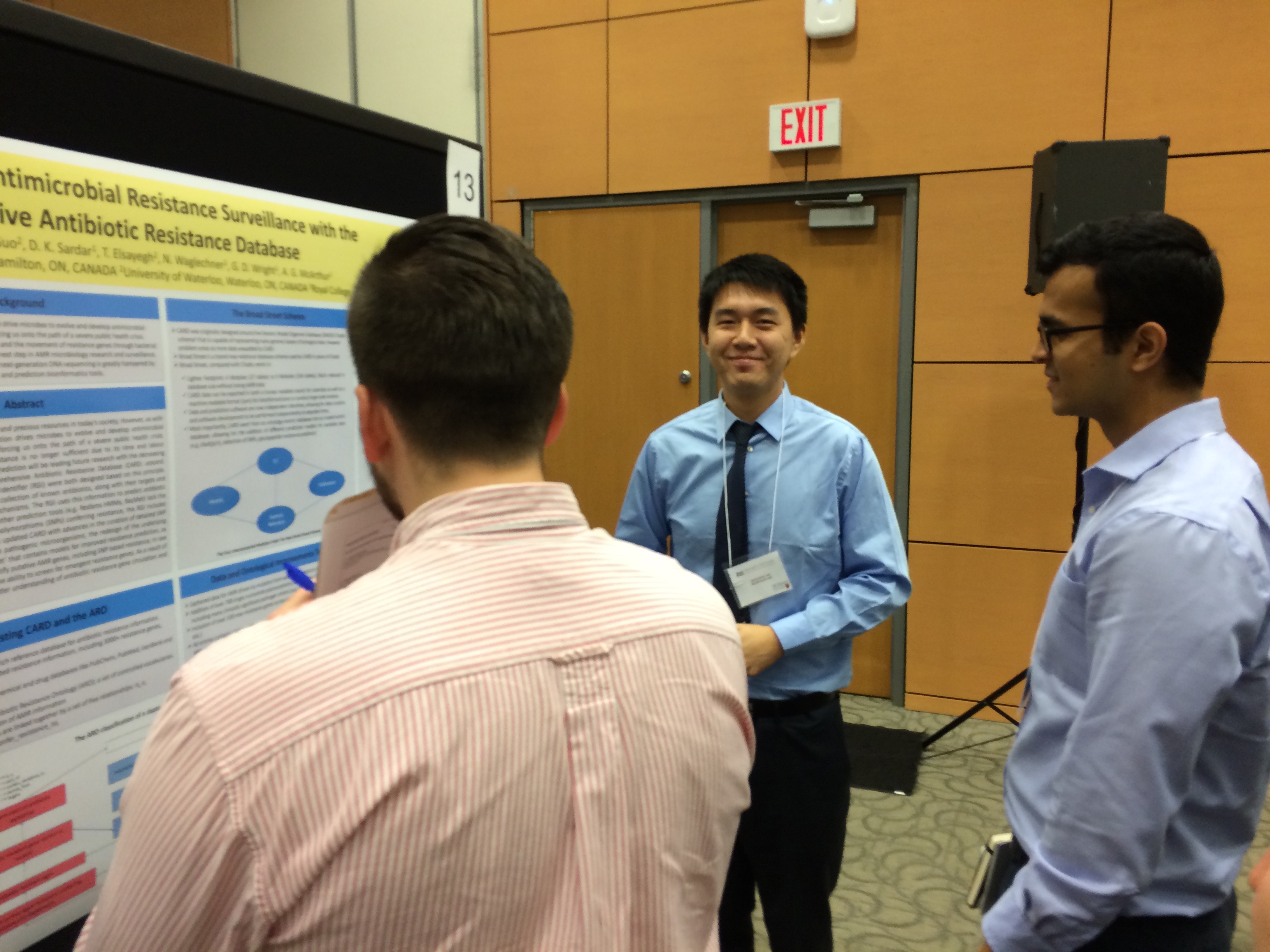

http://traineeday2015.mcmasteriidr.ca

Authors: Freschi et al. Front Microbiol. 2015 Sep 29;6:1036. The International Pseudomonas aeruginosa Consortium is sequencing over 1000 genomes and building an analysis pipeline for the study

Authors: Graham CF, Glenn TC, McArthur AG, Boreham DR, Kieran T, Lance S, Manzon RG, Martino JA, Pierson T, Rogers SM, Wilson JY, Somers CM. Mol Ecol

The McArthur lab is proud to collaborate with colleagues in the Faculty of Science on the metabolic and transcriptional responses to human inactivity and aging under
